Editor’s note: Claims substantiation and a drive for innovative products, among other factors, constantly expand and expose cosmetic scientists to new disciplines—be it in their education, or via the addition of new experts. The present article is a perfect example. It demonstrates how an understanding of molecular biology can be used to test the effects of personal care products on skin’s microbes.
The skin’s rich and diverse community of microorganisms—i.e., the microbiota and its corresponding genetic material, the microbiome—is an integral component of normal human skin. To gain insights into its composition and activities, traditional culture-based techniques have been used but these can be misinterpreted due to the domination of faster-growing organisms. Such methods are now bypassed by tools and techniques supported by advances in DNA amplification, sequencing technologies and new computational tools.
Several studies have inventoried the composition of bacterial communities of the surface of healthy human skin under static conditions. Propionibacterium and Corynebacterium (Actinobacteria) are the dominant organisms in sebaceous areas; Staphylococcus and Streptococcus (Firmicutes) in moist areas; and Acinetobacter (Proteobacteria) in dry sites.1, 2 Furthermore, Grice and Segre3 have addressed endogenous and exogenous host factors that determine colonization.
Nevertheless, the investigation of skin microbiota under the influence of environmental factors such as cosmetics and from a dynamic perspective has as yet been poorly investigated, with only one study targeting the influence of makeup on bacteria diversity.4 Therefore, the aim of the present research was to explore and select appropriate metagenomic tools to investigate the impact of a cosmetic formulation on the skin’s microbial community. Techniques including Automated Ribosomal Intergenic Spacer Analysis (ARISA) and 16S rRNA gene sequence analysis are described here, which can measure ribosome spacer length and sequence polymorphisms, respectively. These techniques were used to process DNA samples taken from a subject using a test and placebo formula, to determine the effects of both formulas on the skin’s microbiome.
Materials and Methods
Test formulas and protocol: A protocol was developed for applying two cosmetic formulas to a healthy, 37-year-old female Caucasian volunteer over the course of 28 days. The volunteer had no symptoms of skin disease and did not use other cosmetic products on her face; she also received no antibiotic treatments during the test period.
The test formula contained a Haberlea rhodopensis plant extract, rich in the polyphenol glycoside myconoside. The placebo formula was identical except without the plant extract.
The test formula was applied daily to the right side of the forehead while the placebo was applied to the left side of the forehead. Each day during the application phase, at a fixed time in the morning and evening, the volunteer applied the same quantity of placebo and test formula to the dedicated zones. The volunteer applied the formulas until completely absorbed to a clean and dry face using gloved fingers.
Skin microbiota sampling: Non-invasive skin samples were collected from each treated zone at fixed dates in the late morning, i.e., 11:00 A.M.–12:00 P.M., by swabbing the application areas with sterile, 5 × 5-cm gauze pre-moistened with a sterile solution of 0.15 M NaCl and 0.1% polysorbate 20. Gauze samples were collected in a 50-mL sterile plastic tube and frozen at -20°C until DNA extraction.
DNA extraction: Each sample tube was fully filled with NaCl and polysorbate 20 solution and horizontally shaken for 15 min at 800 rpm. The NaCl and polysorbate 20 suspension was transferred to new tubes and the gauzes were spin-dryed under sterile conditions to collect most of the suspension volume. Suspensions were then centrifuged at 8,000 rpm for 30 min to obtain a cell pellet.
DNA extraction then was performed based on physical lysis. Briefly, the cell pellet was re-suspended with a hexadecyltrimethylammonium bromide (CTAB) extraction buffer (0.5 mL) and split in two equal parts for duplicate DNA extractions. An equal volume of phenol-chloroform-isoamyl alcohol 25:24:1 was added to each replicate cell suspension. Cells were lysed for 30 sec at 5.5 m/sa, and samples were centrifuged for 10 min at 4°C and 11,000 rpm. The aqueous phase was collected and DNA precipitation was achieved with two volumes of ethanol 100% and 1/10 volume of 5 M NaCl at 4°C, overnight. Finally, purificationb was performed to obtain high purity DNA.
Spacer Length: The ARISA Technique
To analyze microbial diversity and compare microbial communities, ARISA was performed. This technique is based on the analysis of length polymorphisms of the non-codant DNA region (the intergenic spacer) between two highly conserved genes, the 16S rRNA gene and the 23S rRNA gene. The length of this intergenic spacer is taxa-specific. This was accomplished by polymerase chain reaction (PCR) using primers 5ʹ-TGCGGCTGGATCCCCTCCTT-3ʹ (forward) and 5ʹ-CCGGGTTTCCCCATTCGG-3ʹ (reverse), as proposed by Ranjard et al.5
For the PCR mix, 2 µL of DNA was mixed with 1.25 µL of reverse and forward primers (10 µM) and 20.5 µL of distilled water (DH2O). PCR cycles consisted of 95°C for 10 min then 30 cycles of 95°C for 30 sec; 55°C for 30 sec; and 72°C for 1 min followed by 72°C for 15 min with a thermocyclerc. One microliter of the PCR mix was then loaded into a DNA “lab on a chipd,” electropherograms were analyzed, and data was normalized using a bioanalyzere.
This approach generates RISA fragments from most of the dominant bacteria in an environmental sample, which are then profiled. However, the detected peaks often are the result of different microbes possessing spacer regions of identical length, which underestimates the diversity of the microbes that are present.
Order Matters: 16S rRNA Gene Sequence Analysis
To gain further detail on changes to the diversity of microbial communities, 16S rRNA gene sequence analysis was employed. This approach uses the pyrosequencing method of DNA sequencing, which detects pyrophosphate release upon nucleotide incorporation, rather than chain termination with dideoxynucleotides. This analysis was performed by a third partyf and targeted two 16S variable regions: the V1V3 region (16S-0027F forward primer 5ʹ-AGAGTTTGATCCTGGCTCAG-3ʹ and 16S-0533R reverse primer 5ʹ-TTACCGCGGCTGCTGGCAC-3ʹ), producing ~ 520 base pair (bp) amplicons; and the V4V6 region (16S-0515F forward primer 5ʹ-TGYCAGCMGCCGCGGTA-3ʹ and 16S-1061R reverse primer 5ʹ-TCACGRCACGAGCTGACG-3ʹ), producing ~560 bp amplicons.
ARISA could highlight differences in microbial communities without the need for taxonomic identification.
To enable the simultaneous analysis of multiple samples, i.e., multiplexing, multiplex identifiersg were tailed to each end of the primers. PCRs were carried out as follows: 3 ng of each template DNA was mixed with 2.5 µL of each reverse and forward primers (10 µM), 1.25 unit of Taq polymeraseh and 7.9 µL of distilled water (DH2O). PCR cycles consisted of 95°C for 3 min, then 25 cycles of 95°C for 1 min, 58°C for 30 sec, and 72°C for 1 min, followed by a final extension at 72°C for 5 min using a thermocyclerj. Negative controls were included in all steps to check for contamination.
The bioinformatic treatment of raw data sequences was implemented by the same third partyf, which included:
- demultiplexing sequenced samples;
- assembling forward and reverse sequences
(clustering step); - Blastn analysis, i.e., searching nucleotide
databases using nucleotide queries of both clustered and single reads against a curated copy of the 16S RDP database;6 and - making taxonomic assignments by comparing
taxonomy and scores of the 25 best hits for each blasted sequence.7
Whole Genome Sequencing Challenge
The results of 16S taxonomic assignments rely on a PCR amplification, which may induce technical bias. Thus, whole genome sequencing (WGS) also was implemented on the same DNA extracts, in order to challenge initial results. For the library preparation, 25 nanograms (ng) of metagenomic DNA was fragmentedk into an average size of approximatively 250 bp, per the supplier-suggested protocol. Fragmented DNA was then used to synthesize indexed sequencing libraries using a nano-DNA sample preparation kitm also according to the manufacturer. Cluster generation was performedn and libraries were sequencedp for paired-end sequencing with read lengths of 150 base pairs (300 cycles). High throughput sequencing reads were then quality filteredq and only those reads with a quality score of higher than 17 for at least 80% of the read length—i.e., having a probability of correct base calling at approximately 98%—were retained.
Gene catalogs for each sample were created using the MOCAT pipeline.8 Briefly, this pipeline performs quality control on the raw sequencing data reads and removes human contamination by mapping it to the reference human genome. It also assembles the reads and predicts protein-coding genes on the assembled overlapping reads (contigs) and scaftigs—i.e., contigs that were extended and linked using the paired-end information of sequencing reads.
The predicted proteins were then compared to the non-redundant National Center for Biotechnology Information (NCBI) RefSeq database using the Basic Local Alignment Search Tool (BLAST) algorithm for comparing primary biological sequence information.9 Taxonomic analysis was performed based on the NCBI taxonomy, and functional analysis was performed by the MEGAN4 metagenome analyzer using the SEED classification for comparative genomic environments.7, 10, 11
Taxonomic analysis was performed by placing each sequence read onto a node of the NCBI taxonomy, based on gene content. For each read that matches the sequence of some gene, the program places the read onto the lowest common ancestor (LCA) node of those species in the taxonomy that are known to have that gene. This produces what is referred to as an LCA algorithm.
ARISA Method Results
Fingerprints of the extraction replicate samples (see Figure 1a) show a highly similar profile with a perfect superposition of both profiles, i.e., red and dark electropherograms. This demonstrates the high reproducibility of the method.
Comparing the fingerprint of the microbiota corresponding to the right part of the forehead (RFH) with that of the left part of the forehead (LFH), one day before applying the creams (see Figure 1b), a highly conserved profile was observed—i.e., the same peak species (from A to G) were observed but with slight height variations for some. These initial lateralized weak differences justified considering the right and left evolution of fingerprints separately across time and after the application of formulas.
The evolution of the microbiota fingerprint pattern over time on the H. rhodopensis-treated RFH was examined next (see Figure 2). Although the microbial ARISA pattern remained globally constant over time, a quantitative variation was observed for peak D (see Figure 2a). A transient rise in this peak on the RFH was observed after seven days, followed by a regular decrease in this peak on the RFH after 14 days (data not shown), and finally a return to the superposed peak D after 28 days on both sides of the forehead.
A qualitative evolution also was observed since a new peak, CAA (in green), appeared on both the right and left sides of the forehead at day+28 (see Figure 2b). This also was observed at day+14 (data not shown), suggesting it corresponded with a biological event and not to an artifact.
Considering the number of total peaks and changes in peak relative abundance, only small (but detectable) differences in microbial population compositions were observed. However, this encouraged the further analysis using 16S rRNA gene sequences to gain insights into the taxonomic distribution of the samples.
16S rRNA Gene Sequencing Results
For the entirety of the sequenced samples, the recovery of approximately 580,000 reads satisfied the expected read length of ~500 bp. Figure 3 shows initial bacterial relative abundances at the fifth rank level of the classification (Orders) level on the RFH sample by targeting two different variable regions of the 16S rRNA gene. Targeting these two regions clearly gives two different taxonomic highlights of the microbial community: the V4V6 16S rRNA gene region allowed access to a greater diversity, whereas the distribution based on the V1V3 region was heavily dominated by the Actinomycetales.
The V4V6 region was used for additional analysis. Considering the most abundant taxonomic groups, at the Order level, from the RFH and LFH one day before application, a clear lateralization was observed (see Figure 4). Indeed, the Actinobacteridae and Bacillales were more represented on the right side than on the left side.
As observed with the RISA results, the initial lateralized differences between LFH and RFH bacteria diversity required their separate consideration over time, i.e., before and after the application of formulas. Thus, for each forehead zone, microbial sampling was carried out seven, 14 and 28 days. The bacterial diversity for each sample was measured by operational taxonomic units (OTU)—i.e., the terminal node or lowest possible rank in the phylogenetic analysis, specific for each read—as a function of the number of reads (see Figure 5).
For both the RFH and LFH, a strong decrease in bacterial diversity was observed after 14 and 28 days of formula application. However, it must be underlined that after seven days of formula application, the microbial communities were less impacted on the right side, i.e., that treated with the H. rhodopensis-containing formula, than the placebo-treated left side.
WGS Sequencing Results
As noted, WGS was implemented to challenge the results of the 16S-based taxonomic assignments. This sequencing runp recovered 391,866,608 paired reads of 100 bp for the whole of sequenced samples. Comparing the sequence data generated by WGS with the public databases enabled both the taxonomic and functional assignment of these sequences. Figure 6 shows the initial bacterial relative abundances at the Order level on the RFH sample.
WGS confirmed the previously observed lateralized difference between the left and right forehead sides at the initial collecting time, i.e., one day before application (data not shown). Furthermore, the taxonomic assignation of sequences through the WGS approach not only confirmed that Bacillales and Actinomycetales were the major bacterial orders constituting the forehead microbiota, but also enabled the detection of eukaryote representatives such as Malasseziales fungi.
The ARISA profiles also showed low complexity, which suggests unbalanced bacterial consortia with highly dominant bacterial members.
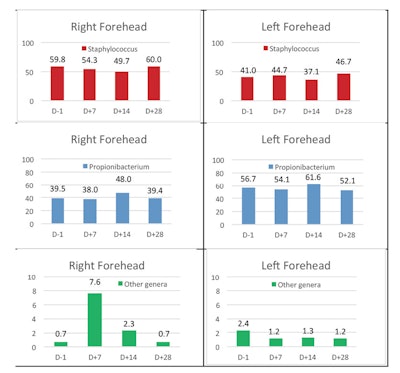
The global examination also confirmed that bacterial communities of the forehead were strongly unbalanced, with the two highly dominant genera Propionibacterium and Staphylococcus together encompassing 99.3% and 97.7% of the assigned genera on the RFH and LFH, respectively (see Table 1).
Next, the evolution over time of the relative abundance of the sequences assigned to the Staphylococcus genera, the Propionibacterium genera and the others were considered. Fluctuation of the two dominant genera, Staphylococcus and Propionibacterium, over time was observed on both the RFH and LFH, with no clear tendency. The relative abundance of the two also remained stable over time, on both forehead sides.
In contrast, a different effect was observed with the other, minor genera: an increase (from 0.7% to 7.6%) in relative abundance was observed on the right side one day before and seven days after application of the formula containing H. rhodopensis extract; whereas on the left, placebo- treated side, a decrease (from 2.4% to 1.2%) was observed.
Once filtering and assembling the data corresponding to the eight samples (i.e., four samples for both zones), more than 61,100 genes were predicted and their corresponding proteins defined. To identify the samples that showed a significant evolution of functional abundance in the assigned genes, compared with the initial sampling (i.e., one day before application of formulas), a non-parametric Wilcoxon signed-rank test12 was used.
From the all possible sample combinations, the only distinct comparison of populations that could be made was between one day before and 28 days after the application of the H. rhodopensis test formula (RFH). Here, the p value of 1.47e-05 < 0.05–0.05 was the upper threshold value. On the placebo-treated LFH, between one day before and 28 days after application, populations could not be considered distinct (p value 0.7982 > 0.05); a global view is given in Figure 7.
Examining this chart shows a relatively stable functional status of the skin microbiota following the application of the H. rhodopensis-containing formula. A deeper look into the evolution of the functional subsystems also shows some evolution in proteins between day-1 and day 28. Nevertheless, it must be noted that the identification of functional subsystems showing a significant evolution over time would require statistical analyses with biological replicates of data from several volunteers.
ARISA Delivers, to an Extent
As stated, healthy human skin microflora is still poorly understood, as it is diverse and difficult to analyze using traditional methods. It becomes even more challenging to study and understand its evolution during the application of a cosmetic formula. Thus, the first objective was to evaluate whether the ARISA fingerprint method could detect potential modifications to skin bacterial communities following the application of cosmetic formulas.
Indeed, yes, in general, the technique successfully produced a community fingerprint based on length polymorphisms. According to the test facilityf, it also was rapid, required low amounts of DNA, and allowed for the simultaneous analysis of multiple samples. Furthermore, it could highlight effects on or differences between microbial communities without the need for direct taxonomic identitification.
The ARISA approach differs from manual methods in that PCR is conducted with a fluorescence-tagged oligonucleotide primer, providing laser detection of fluorescent DNA fragments. Furthermore, ARISA-PCR may generate DNA fragments of up to 1,400 bp in length, although it must be emphasized that the vast majority of the intergenic spacer lengths fall between 150 bp and 600 bp, with mean spacer lengths seemingly higher for Gram-positive than Gram-negative microbes.13
Important to note is the fact that the ARISA fingerprinting technique enables single organisms to contribute more than one peak to an ARISA profile. And that unrelated microorganisms may possess spacer regions of identical lengths, which are represented in the ARISA profile by a single peak, underestimating microbe diversity.
The ARISA profiles also showed low complexity, which suggests unbalanced bacterial consortia with highly dominant bacterial members. The ARISA pattern also remained stable over time, with only a slight evolution in amplification profiles. From this, it may be assumed that the test formulas did not have detrimental effects on the native bacteria communities, and that the constancy of ARISA profiles was a result of a moderate impact by the formulas.
However, the ARISA fingerprint method does not establish links between the observed peak profiles and the underlying microbial community. This is because each peak is the result of multiple amplicons sharing the same length, most likely from different organisms.
16S Sequencing Takes it Further
To shed additional light on the composition of bacterial communities, 16S-amplicon based sequencing was implemented. As expected, this approach proved to be more informative than ARISA. Sequencing by 16S confirmed the forehead bacterial community was unbalanced and highly dominated by Actinobacteridae, mainly represented by the Propionibacterium genera, and Bacillales, mainly represented by the Staphylococcus genera.
Comparing the taxonomic distributions produced with two different targeted 16S variable regions clearly demonstrated that the amplicon-based sequencing strategy did not enable the absolute elucidation of the bacterial community composition. V1V3 region amplification favored the Actinobacteridae and displayed a forehead skin community highly dominated by Propionibacterium. In contrast, V4V6 amplification allowed a best representation of other Orders, including the Bacillales (Staphylococcus).
The choice of the 16S variable region and primer sets used to produce 16S amplicons can therefore greatly affect the final taxonomic description of bacterial communities. This amplicon-based sequencing approach is then fully adapted for the relative comparison of samples, or for the monitoring of taxa evolution over time, but it does not allow the absolute and quantitative assessment of the skin bacterial community. Moreover, highlighting the taxonomic composition through the sequencing of only 16S rRNA genes does not provide information about the functional status of the bacterial community—i.e., it tells you who is there but not who can do what.
WGS Goes the Distance and Beyond
The study was therefore extended using WGS to more deeply define the full genetic diversity and gain insights about the gene functions associated with the collected skin microbiota. Direct sequencing enables both taxonomic and functional assignments of assembled sequences. This approach also does not rely on the prior PCR amplification of a single biomarker gene, which is a possible source of bias. It is therefore accepted that direct sequencing enables a less biased description of the bacterial community.
However, this approach has two constraints. First, the whole genome sequencing will mainly address the most dominant taxa of the bacterial community. Treatments that would only affect some rare taxa of the microbiota might hardly be detected using this approach. In fact, in the present study, more than 95% of the overall bacteria fell into only two genera: Staphylococcus and Propionibacterium.
Differences were only observed on the RFH between D-1 and D+28 days during the application of H. rhodopensis; the differences were therefore attributed to the extract.
Second, since the whole of genome sequencing addresses overall genes, a consolidated sequencing study requires a much higher sequencing effort, in terms of the number of sequence reads to produce—i.e., sequencing depth; and the statistical bioanalysis required on an extended panel of biological replicates—i.e., volunteers cohorts, sampling points, etc. Yet, since direct sequencing targets the overall genes from the microbiota, it enables the detection of sequences from other skin microorganisms such as fungi that cannot be accessed by the 16S bacterial sequencing approach.
Here, besides the general observation that the application of a cosmetic formula with conventional additives affects the corresponding microbiome, trends were sought potentially related solely to the presence of H. rhodopensis. In doing so, the 26 general SEED functional subsystems (see Figure 7) were considered.
As mentioned, significant differences only were observed on the RFH between D-1 and D+28 days during the application of a formula containing H. rhodopensis. It thus could be suggested that the differences were related to the presence of the extract.
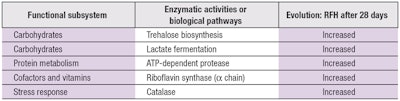
More precisely, among the 26 general SEED functional subsystems considered, those impacted by the extract included:
- Carbohydrate transport, synthesis and degradation;
- Protein metabolism;
- Cofactor and vitamin transport, synthesis
and assimilation; - Cell wall and capsule biosynthesis;
- Fatty acid, lipid and isoprenoid biosynthesis;
- Respiration; and
- Stress response.
Examples of enzymatic activities or biological pathways that were found in higher or lower normalized abundances are shown in Table 2.
Conclusion
These studies show that several methodologies are available for the characterization of skin microbiota and the monitoring of its evolution over time. The ARISA fingerprint method was confirmed, albeit less accurate, as a useful first attempt for the preliminary screening of numerous samples to decide which samples warranted deeper analysis.
The 16S-based taxonomic characterization was found to provide much more indication as to which bacteria are present in a given environment, and how their relative distribution may differentially evolve under an applied treatment and/or over time. This second methodology could easily be applied using a very large number of samples. However, it may suffer from technical bias, which is why a third methodology, WGS, was deemed suited to contribute to both taxonomic and functional characterization of the skin microbiota.
This paper on the first dynamic metagenomic study provides valuable methodological guidance about how to gain a better understanding of the microbiota profile of healthy skin over time. It serves as a starting point to develop a deeper characterization of the relationships between cosmetic ingredients, skin microbiota and cutaneous epithelium.
References
- Gao et al, Molecular analysis of human forearm superficial skin bacterial biota, PNAS 104(8) 2927-2932 (2007)
- Z Gao et al, Quantitation of major human cutaneous bacterial and fungal populations, J Clinic Microbiol 48(10) 3575-3581 (2010)
- EA Grice and JA Segré, The skin microbiome, Nat Rev Microbiol 9(4) 244-253 (2011)
- T Staudinger, Molecular analysis of the prevalent microbiota of human male and female forehead skin compared to forearm skin and the influence of make-up, J Appl Microbiol 110(6) 1381-1389 (2011)
- L Ranjard et al, Sequencing bands of ribosomal intergenic spacer analysis fingerprints for characterization and microscale distribution of soil bacterium populations responding to mercury spiking, Appl Environ Microbiol 66 5334–5339 (2000)
- JR Cole et al, Ribosomal Database Project: Data and tools for high throughput rRNA analysis, Nucl Acids Res 42 (database issue) D633-D642 (2014)
- DH Huson et al, MEGAN analysis of metagenomic data, Genome Res 17(3) 377-86 (2007)
- JR Kultima et al, MOCAT: A metagenomics assembly and gene prediction toolkit, PLoS One 7(10) e47656 (2012)
- SF Altschul et al, Gapped BLAST and PSI-BLAST: A new generation of protein database search programs, Nucleic Acids Res 25(17) 3389-402 (1997)
- DH Huson et al, Integrative analysis of environmental sequences using MEGAN4, Genome Res 21(9) 1552-60 (2011)
- R Overbeek et al, The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes, Nucleic Acids Res 33(17) 5691-702 (2005)
- F Wilcoxon, Individual comparisons by ranking methods, Biometrics Bull 1(6) 80-83 (1945)
- MM Fisher and EW Triplett, Automated approach for ribosomal Intergenic spacer analysis of microbial diversity and its application to freshwater bacterial communities, Appl Environ Microbiol 65(10) 4630–4636 (1999)