Skin represents the first line of defense against various stresses including pathogens and injuries. Within skin, cutaneous homeostasis is maintained by permanent cross-talk between skin cells, epidermal keratinocytes in particular, and cells of the immune system residing in the skin or recruited through the local production of cytokines and chemokines (see Figure 1).
The antigen-presenting cells—dendritic cells (DC) or Langerhans cells (LC)—are major early actors in the inflammatory or allergic response.1, 2 Recruitment and activation of immune or blood cells, such as T lymphocytes (TL), polymorphonuclear leukocytes (PMN) or others, leads to keratinocyte stimulation and the installation of an inflammatory loop. This inflammatory loop represents the innate defense of the skin but its deregulation may lead to more or less severe skin symptoms.
Conventional and recently developed anti-inflammatory therapies mainly target immune cells but are neither optimal nor comfortable for patients. Therapies that employ anti-inflammatory/immuno-suppressive drugs, e.g., corticoids, and cyclosporine/tacrolimus drugs have side effects and these materials are not fully efficient because they target only subsets of inflammatory cells. Novel therapies use biotechnology-derived products such as humanized monoclonal antibodies but these proteins require injection. It is therefore important to find alternative or complementary strategies to better control inflammation; for instance, by blocking either inflammation at the keratinocyte level, or the capacity of immune cell recruitment and epidermal hyperplasia (see Figure 2). Topical formulations of active ingredients are also interesting for limiting or preventing the effects of environmental stress or aging.
Leucocyte infiltrates produce pro-inflammatory cytokines that act synergistically on keratinocytes to induce the production of innate immunity molecules, such as antimicrobial peptides (defensin, S100 proteins, elafin, LL37, etc.), scavenger proteins (heme-oxygenase 1) and chemokines (CXCL8).3 It should be noted that a number of antimicrobial peptides exhibit chemo-attractive activity and that, conversely, chemokines can be antimicrobial. These systems cooperate for the maintenance and evolution of the inflammatory pathology, e.g. psoriasis. This physiopathology relates to strong modifications of the epidermis including hyperplasia and differentiation loss, and results in a modification of the expression of a number of keratinocyte structural markers.
Modeling Skin Inflammation: Cytokines and the Keratinocyte
Several markers of skin inflammation have been described and keratinocyte responses to many individual cytokines have been reported.4, 5 However, no clear and comparative profiling of keratinocyte response has been shown to date. In addition, infiltrating leukocytes and especially TL produce many different cytokines that have potential to synergize, or in some cases antagonize, to induce specific responses.6-8 Thus, modelling the inflammatory pathology in keratinocytes in vitro was initiated by the authors via two parallel strategies performed on normal human epidermal keratinocyte (NHEK) monolayer cultures (see Figure 3). The final step was to associate some of these cytokines to reproduce the profile obtained with activated TL.
TL-challenged NHEK cultures: In the first approach, NHEK cultures were challenged with culture supernatants from activated TL isolated from psoriatic biopsies to identify keratinocyte biomarkers of this inflammation. The first step was to isolate and amplify TL lines from psoriasis lesions6, 8 and to prepare culture supernatants of activated TL (anti-CD3/anti-CD28 activation) keratinocyte cultures. NHEK cultures were then challenged with these psoriatic T cell-derived supernatants to identify the modification of the keratinocyte transcriptional profile, and to select a restricted set of highly modified markers as tracers of keratinocyte inflammation in subsequent modeling or pharmacological experiments (see Figure 4). Non-activated LT media were used as a control and the red dots in the figure correspond to significantly up-regulated genes; green dots to down-regulated genes.
As expected, the transcription of genes coded for antimicrobial peptides, chemotactic molecules and epidermis structure/differentiation antigens were strongly modulated, and a representative set of the four most dramatically modified markers—S100A7, HBD2, CXCL8 and KRT10—was selected as reference genes for subsequent studies and model design.
Cytokines and inflammation: In parallel, a second approach involved the systematic screening of more than 40 cytokines and growth factors in NHEK cultures (see Figure 4 and Figure 5) to identify keratinocyte-responding cytokines, with special emphasis on the four previously selected markers that were found to characterize the inflammatory response of the keratinocyte. The cytokines that modified the expression of at least one gene by at least a fivefold increase or decrease are highlighted.
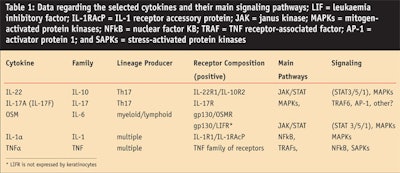
After quantitative reverse transcription polymerase chain reaction (RT-qPCR) confirmations and comparisons, the authors selected a set of five cytokines that induced a clear inflammatory response in keratinocytes and that could be taken as candidates for a synergistic pro-inflammatory challenge. These cytokines were IL-22, IL-17, oncostatin M (OSM), IL-1alpha (IL-1α), and tumor necrosis factor alpha (TNFα).7, 9
Each of these cytokines belong to a specific family and signal via different receptors and pathways (see Table 1). These cytokines were significantly produced by activated TL derived from psoriasic skin lesions (not shown). As expected, the association of these cytokines led to strong keratinocyte responses (see Figure 6), more so than those obtained with the conditioned media from activated TL.
As an illustration of this synergistic activity, the authors showed that the mix of five cytokines in NHEK cultures reproducibly up-regulated the expression of S100A7 and hBD2—1,000 fold and 50,000 fold above unstimulated NHEK, respectively—and down-regulated the expression of KRT10, tenfold below unstimulated NHEK (data not shown). Comparatively, the effects of individual cytokines were much lower (see Figure 6).
After testing various associations of 2–4 of cytokines and associations with other unrelated cytokines and growth factors, the authors determined that a minimal synergistic association of OSM + IL-17 +TNFα (or IL-1)10 was necessary for a nearly optimal pro-inflammatory response (see Figure 6). These results indicate that optimal synergy engages at least one cytokine that signalizes through the JAK/STAT pathway, especially STAT3,11 and requires cooperation with inducers of NFkB and MAPK pathways; IL-17 is also crucial for this synergy. A more detailed analysis of the implication of these signaling pathways in the keratinocyte response is under investigation.
Applications and Testing
The described studies were aimed at in vitro modeling of psoriasis using adequate stimulations and validated analytic systems. This work then led to the design of a series of models for the research and study of compounds that could modulate this inflammatory profile. Figure 7 summarizes the signaling of inflammatory cytokines in NHEK and the type of assays or analyses that are currently performed in this model.
As was shown earlier, NHEK stimulation can be analyzed at the mRNA level using RT-qPCR and DNA arrays. RT-qPCR arrays (qPCR arrays) are the method of choice for a direct multiplex analysis of sets of markers; i.e., genes coded for antimicrobial peptides, chemokines, keratinocyte differentiation markers and other markers of interest. Such methods give a good indication of which group of genes is targeted by a product, and whether the observed effects return to a basal condition. In parallel, the modifications can be studied at the protein level by quantitative immunofluorescence in the NHEK cell layers, i.e., immunolabeling of S100A7 and KRT10, and by ELISA or related assays in culture media, i.e., ELISA of hBD2 and CXCL8 (see Figure 7).
After receptor binding, the cytokines induce specific signaling including STAT-3 phosphorylation, which is measured by Western blot and immunofluorescence; MAPKs phosphorylation, measured by Western blot of phospho-ERK, phospho-P38 and phospho-JNK; and NFkB nuclear translocation, measured by immunofluorescence (see Figure 7). This synergistic signaling induces a profound and specific modification of keratinocyte transcriptome that mimics the one induced by activated TL derived from psoriasic skin lesions. The target marker’s skin innate immunity and epidermal differentiation can be measured at the mRNA level by real-time PCR, and at the protein level by specific immunodetection methods such as immunofluorescence and ELISA.
All of these approaches can be used for researching compounds that could modulate skin inflammation status. In terms of compound testing, ELISA assays for soluble factors and immuno-fluorescence using an automated microplate system are well-adapted to the screening of compounds in cellular assays (commonly called high-content screening). Screening methods may directly analyze the effects on target proteins but also may analyze early signaling events (phosphorylations and nuclear translocations) that occur in less than 30 min. The authors have tested reference compounds on these models that indicate that some JAK/STAT inhibitors almost completely reverse the keratinocyte inflammatory phenotype. In contrast, commonly used anti-inflammatory compounds such as cyclosporin A or glucocorticoids had no effect (data not shown).
Conclusion
The keratinocytes and overall epidermis are considered as a protective structure that directly responds to only few external stimuli, leading to rather low quantitative responses. This could explain why the cell structures are well-protected against external aggressions and why non-pathological and non-lesional skin is not inflammatory under acceptable living conditions. However, under specific stimulations mainly mediated by immune cells, the keratinocyte can exhibit strong responses thus becoming a major actor in the innate immunity of the skin. Under these pathological conditions, keratinocytes produce high amounts of chemo-attractant molecules, hence leading to the installation of an inflammatory loop and chronic skin inflammation, in the case of psoriasis.
Here, the authors identified inflammatory cytokine combinations that can stimulate keratinocytes, mimicking the strong response induced by inflammatory TL infiltrates. In these mixes, Th17 cytokines are of major importance, as has been described in numerous publications as well as the authors’ models.
This work has led to the design of proprietary models for the evaluation of products, drugs or conditions. The knowledge of signaling pathways induced by the cytokines engaged in this process may orient the research of new anti-inflammatory molecules that are able to block the recruitment of immune cells and reverse the inflammatory/lesional status of the skin. In pathological conditions, such keratinocyte targeting molecules could optimize the effects of the conventional therapies targeting immune cells. In soft inflammatory conditions or in protective formulations, topically applied actives would complete a panel of health care solutions.
References
1. D von Bubnoff, E Geiger and T Bieber, Antigen-presenting cells in allergy, J Allergy Clin Immunol 108 329–339 (2001)
2. RM Steinman and J Banchereau, Taking dendritic cells into medicine, Nature 449 419–426 (2007)
3. AS Büchau and RL Gallo, Innate immunity and antimicrobial defense systems in psoriasis, Clin Dermatol 25 616–624 (2007)
4. JC Lecron, F Morel, K Boniface and FX Bernard, Keratinocytes as targets for cytokines in skin inflammation, in: Recent Advances in Immunology of the Skin, S Saeland ed, in press (2009)
5. K Boniface et al, Keratinocytes as targets for IL-10-related cytokines: A putative role in psoriasis pathogenesis, Eur Cytokine Netw 16 309–319 (2005a)
6. K Boniface et al, A role for T cell-derived Interleukin 22 in psoriatic skin inflammation, Clin Exp Immunol 150 407–415 (2007a)
7. K Boniface et al, Oncostatin M secreted by skin infiltrating T lymphocytes is a potent keratinocyte activator involved in skin inflammation, J Immunol 178 4615–4622 (2007b)
8. NJ Wilson et al, Development, cytokine profile and function of human interleukin 17-producing helper T cells, Nat Immunol 8 950–957 (2007)
9. K Boniface, FX Bernard, M Garcia, AL Gurney, JC Lecron and F Morel, IL-22 inhibits epidermal differentiation and induces proinflammatory gene expression and migration of human keratinocytes, J Immunol 174 3695–3702 (2005b)
10. Patent PCT/EP2005/014198
11. S Sano et al, Stat3 links activated keratinocytes and immunocytes required for development of psoriasis in a novel transgenic mouse model, Nat Med 11 43–49(2005)